|
4/19/2023 0 Comments Logistic regression r studio Plot.data <- gather(plot.data, key=group, value=prob, a:c) Plot.data <- ame(a=a_probs, b=b_probs, c=c_probs, X1=X1_range) # first you have to get the information into a long dataframe, which is what ggplot likes :) Or, you can do it in ggplot2! library(ggplot2) library(tidyr) # Add the line for people who are in the c group # Add the line for people who are in the b group Xlab="X1", ylab="P(outcome)", main="Probability of super important outcome") Plot time! # We'll start by plotting the ref group: # Compute the probibilities (this is what will actually get plotted):Ī_probs <- exp(a_logits)/(1 + exp(a_logits))ī_probs <- exp(b_logits)/(1 + exp(b_logits))Ĭ_probs <- exp(c_logits)/(1 + exp(c_logits)) X2_val <- mean(data$X2) # by plugging in the mean as the value for X2, I'll be generating plots that show the relationship between X1 and the outcome "for someone with an average X2". You make a separate equation for each group by plugging in different values for the group dummy codes. 
You’ll need to plug in values for all but one variable – whichever variable you decided will be displayed on the x-axis of your plot. These equations need to include every coefficient for the model you ran. 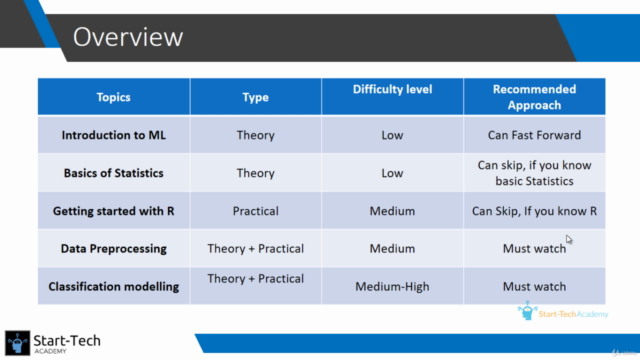
Next, compute the equations for each group in logit terms. (The range we set here will determine the range on the x-axis of the final plot, by the way.) X1_range <- seq(from=min(data$X1), to=max(data$X1), by=.01) That’s the only variable we’ll enter as a whole range. Note: If you were working in SPSS (or for some other reason you have run a model but can’t generate a plot for it), you can enter in your coefficients here, like this: b0 <-0.5872841 # interceptįirst, decide what variable you want on your x-axis. # X1:groupb X1:groupc X2:groupb X2:groupc # Number of Fisher Scoring iterations: 7 model$coef # (Intercept) X1 X2 groupb groupc X1:X2 # Residual deviance: 205.46 on 290 degrees of freedom # Null deviance: 412.88 on 299 degrees of freedom # (Dispersion parameter for binomial family taken to be 1) # note this use of exponent in a formula will give us all 2-way interactions For example contrasts="eatment" will make it traditional dummy coding if it isn't already. # If you want to change what contrasts will run, you can add an argument to the glm() call. And we can see that a is the reference group. # d 0 0 1 # Great, that's what we wanted. Get the coefficients from your logistic regression modelįirst, whenever you’re using a categorical predictor in a model in R (or anywhere else, for that matter), make sure you know how it’s being coded!! For this example, we want it dummy coded (so we can easily plug in 0’s and 1’s to get equations for the different groups). set.seed(24601) # setting this so the random results will be repeatable We’ll run a nice, complicated logistic regresison and then make a plot that highlights a continuous by categorical interaction. The p-value is 0.00000145, p-value is too model and hence the model is statistically significant.Plotting the results of your logistic regression Part 1: Continuous by categorical interaction 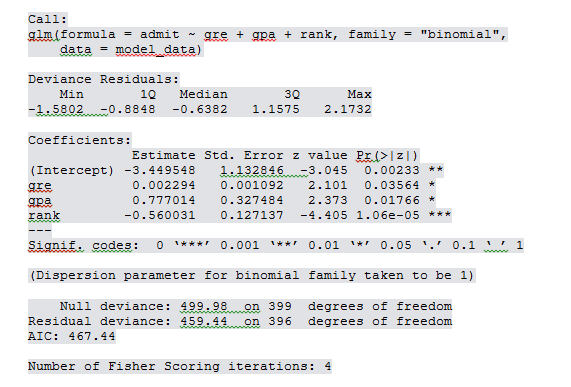
With(mymodel, pchisq(viance – deviance, df.null-df.residual, lower.tail = F)) Logistic regression we can also try for the goodness of fittest. Total 48+5=53 correct classification and 21 misclassifications in test data. Tab2 <- table(Predicted = pred2, Actual = test$admit) Let’s do the prediction based on the above model p1 0.5, 1, 0) Now you can see that gpa pvalue goes down further compared to the previous model. Residual deviance: 371.81 on 320 degrees of freedom 
Let’s drop gre and re-run the model because gre is not significant. In this case gre and level rank 2 is not statistically significant. More stars indicate more statistical significance. Residual deviance: 369.99 on 319 degrees of freedom Null deviance: 404.39 on 324 degrees of freedom (Dispersion parameter for binomial family taken to be 1) As usual create training and test datasets basis 80:20 ratio. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
